Blue Light Can Activate Neurons Without Optogenetic Manipulation
Thiago Arzua is currently a PhD Candidate at the Medical College of Wisconsin, Milwaukee, WI. Follow them on Twitter @thiagoarzua
Optogenetics sounds, on first mention, like something from science fiction—the ability to activate neurons and other cell types with the right wavelength of light. Nevertheless, ever since its inception over a decade ago, optogenetics has been used extensively with an emphasis on electrically excitable cells, such as neurons. Optogenetics has been used to make neurons fire, express specific proteins, and initiate cell-to-cell signaling. Optogenetic manipulation is performed by genetically engineering neurons to express exogenous proteins, such as channelrhodopsins or archaerhodopsins, that are found in light-sensing bacteria. Some of the wavelengths used in optogenetics include 475 nm (blue light), 530 nm (green light), and 612 nm (red light); these wavelengths can activate different variants of channelrhodopsins. Because of the widespread use of optogenetics, scientists are now asking a more basic question: to what extent could light stimulation itself also affect cells? In their eNeuro publication, Tyssowski and Gray examined if, in the absence of any exogenous channelrhodopsins, brain cells would exhibit transcriptional activation upon stimulation with flashing blue, green or red light.
The authors focused on a light exposure duration of 1 or 6 hours, as several optogenetics studies have stimulated neurons for hours or even days. Cortices were isolated from mid-embryonic or neonatal CD1 or C57/Bl6 mice that did not express any channelrhodopsins. Mixed cortical cultures containing dissociated neurons and glia were plated. The cultures were exposed to pulses of light, predominantly at 100% light intensity: 3.9 mW/mm2 (475 nm, blue light), 1.9 mW/mm2 (530 nm, green light) and 2.2 mW/mm2 (620 nm, red light). Control cultures were not stimulated with light and were kept in darkness. After 1 or 6 hours, total RNA was isolated for quantitative PCR analysis of mRNA levels. Fold changes in gene expression levels were determined by comparing cultures exposed separately to each wavelength of light to the non-stimulated control cultures.
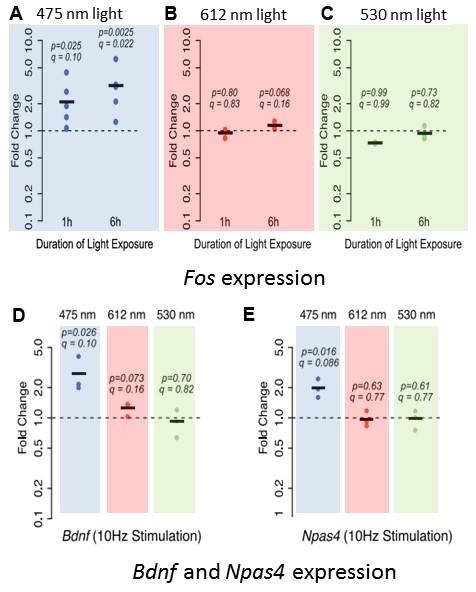
One of the key findings of Tyssowski and Gray’s eNeuro publication is that at the transcriptional level, neurons exposed to blue light were activated, whereas neurons exposed to red or green light were not. Mixed cortical cultures were exposed for 1 or 6 hours to 10 Hz, 2 ms pulses of blue, red or green light. The immediate early gene Fos was used as a marker of neuronal activity. Blue light exposure for 1 or 6 hours increased Fos expression by 2.1- and 3.2-fold, respectively, above control levels (Figure 1A). Cultures exposed to either red or green light, however, had no significant changes in Fos expression at either time point compared to control cultures (Figure 1B, C). Differences in the power used for the three wavelengths of light stimulation were excluded as the cause of the varied transcriptional response. Tyssowski and Gray also tested other neuronal-activity-regulated genes, namely Brain-derived neurotrophic factor (Bdnf) and Neuronal PAS Domain Protein 4 (Npas4). Similar to Fos expression, stimulation with blue light increased the expression of both Bdnf and Npas4 (Figure 1D, E). These results demonstrate that even in the absence of any channelrhodopsin, blue light stimulation alone transcriptionally activates neurons in culture.
The authors also plated neurons on multi-electrode array plates, which enable sensitive measurement of the electrical activity of the neurons. Interestingly, when the authors examined the electrical activity of the neurons exposed to different wavelengths of light, there was no difference in spontaneous action potentials between experimental groups. This finding suggests that blue light is not necessarily depolarizing the membrane of these neurons to increase Fos, Bdnf, or Npas4 expression. However, it is important to point out that there were no changes immediately following blue light exposure, and that prolonged exposure could indeed depolarize the cells.
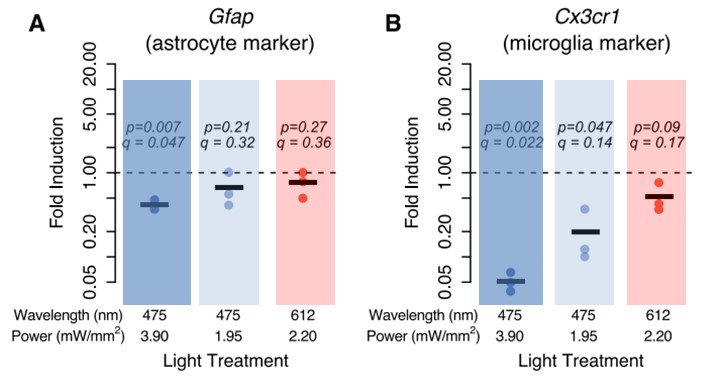
To test if blue light exposure affected any non-neuronal cell types, the authors measured the expression of an astrocyte marker (Gfap) and a microglia marker (Cx3cr1) following 6 hours exposure with either blue or red light. There was a 2.4-fold decrease in Gfap expression in cultures treated with blue light, as well as a significant decrease, up to 20-fold, in Cx3cr1 expression (Figure 2A, B). These changes seemed to be independent of the power output of the light, given that even at 50% power (1.95mW/mm2), blue light still had an effect. These findings suggest that either the expression of the markers was decreased, indicating some change in astrocyte and microglia, or that blue light exposure may have killed these cells.
Overall, Tyssowski and Gray raise some important concerns regarding optogenetics studies. With a large number of experiments relying on optogenetic activation of neurons, it is important to consider whether the exogenous channelrhodopsin is indeed the mediator of that activation. The authors have shown that, at the transcriptional level, blue light stimulation at 475 nm is capable of activating neurons that do not express an exogenous channelrhodopsin. Fos, Bdnf, and Npas4 expression were all significantly upregulated, while the expression of markers for astrocytes (Gfap) and microglia (Cx3cr1) was severely downregulated. There are follow-up studies that can be done to determine the potential mechanisms behind these changes, such as measuring changes in oxidative stress and possibly neuroinflammation. For researchers working on optogenetics, however, the take-home message from this paper is to carefully consider appropriate controls for every experiment, to confirm observations are channelrhodopsin-dependent.
Read the full article:
Blue Light Increases Neuronal Activity-Regulated Gene Expression in the Absence of Optogenetic Proteins
Kelsey M. Tyssowski and Jesse M. Gray
This Reader's Pick was reviewed and edited by eNeuro Features Editor Rosalind S.E. Carney, D.Phil.
FOLLOW US
POPULAR POSTS
TAGS
CATEGORIES


 RSS Feed
RSS Feed




